Evolving Immunity:
A Response to Chapter 6 of Darwin’s Black Box
By Matt Inlay
Draft 1.0 – email author for comments and feedback
![]()
Immune systems are generally characterized by their ability to distinguish between self and non-self cells, tissues, or molecules, and to eliminate the non-self (Janeway 2001). The mammalian immune system is an incredibly complex and intricate system that can recognize non-self and provide protection from a wide variety of pathogens. While there is a high degree of interconnectivity between its components, the immune system can be loosely divided into two subsystems, the innate and the adaptive immune systems. In general, innate immunity is a non-specific, inducible response to pathogens. It is immediate in action, yet short-lived. On the other hand, the adaptive immune system is much more specific, but takes longer to activate. It also features immunological memory, and can augment itself to respond more quickly and with greater specificity to future infections of similar pathogens. Both systems work together to provide protection against a diverse and rapidly-evolving array of pathogens.
While much is known about the evolution of the immune system, the details of their origin remain to be elucidated. The innate immune system is the more ancient of the two systems, with roots deep in the deuterostome branch of the bilaterians, roughly one billion years ago. Conversely, the adaptive immune system appeared more recently and quite suddenly, around 450 million years ago with the emergence of the gnathostomes, more commonly called the jawed vertebrates. Because of its high degree of complexity and interconnectivity, the mammalian immune system has been labeled as "irreducibly complex", and its evolution and origin through "Darwinian" mechanisms challengedi.
Michael Behe, in his book Darwin’s Black Box (Behe 1996), defines irreducible complexity (IC) as:
"[A] single system composed of several well-matched, interacting parts that contribute to the basic function, wherein the removal of any one of the parts causes the system to effectively cease function." (page 39).
Unfortunately, each of the terms "system", "well-matched", "interacting", "parts", and "function" can have multiple meanings, depending on the context, so nonarbitrarily labeling a system as IC is not an easy task. For instance, Behe uses the term "system" differently than the term is used in "immune system". His use of "system" refers specifically to biochemical systems. What is the difference? The components, or parts of a biochemical system are molecular in nature, like proteins or complexes of molecules. "System" as is used in immune system refers not just to molecules, but to the cells and tissues involved in conferring immunity. The immune system as a whole contains numerous biochemical systems within it. It is important to note this distinction because Behe only tries to categorize biochemical systems as IC.
Systems, whether biochemical or not, are often defined according to their function. However, the term "function" is also nebulous. The "function" of the immune system is rather broad, encompassing any action involved in either immune recognition or response. Even the skin is considered to be part of the immune system. The "function" of a biochemical system is much more specific. Here, function refers to a specific biochemical function, such as the formation of a blood-clot, the spinning of a flagella to produce locomotion, or the synthesis of a biomolecule from a precursor. This type of function usually consists of a series of biochemical reactions. This is important because Behe is trying to make a specific argument about the origin and evolution of IC biochemical systems.
Immediately after defining IC, Behe states:
"An irreducibly complex system cannot be produced directly (that is, by continuously improving the initial function, which continues to work by the same mechanism) by slight, successive modifications of a precursor system, because any precursor to an irreducibly complex system that is missing a part is by definition nonfunctional. An irreducibly complex biological system, if there is such a thing, would be a powerful challenge to Darwinian evolution. Since natural selection can only choose systems that are already working, then if a biological system cannot be produced gradually it would have to arise as an integrated unit, in one fell swoop, for natural selection to have anything to act on."
Behe is trying to suggest that because of the nature of IC, biochemical systems that possess this quality could not have evolved through random mutation and natural selection, or "blind-watchmaking". Essentially, Behe is using IC to disprove evolution. As an alternative, Behe suggests that the only way that IC biochemical systems could have originated is through intelligent intervention. He offers as evidence the fact that all IC systems whose origins are known were designed by intelligent agents, us.
There are many problems with Behe’s argument. Basically, Behe is saying that because we do not know how an IC system could have evolved, it didn’t evolve. This type of argumentation is referred to as an argument from ignorance. The problem with arguments from ignorance is that it makes a conclusion based on a lack of knowledge. There is still much we do not know about evolution. What seems like a conundrum right now may be easily resolved in the future. Biology, and science in general, is filled with mysteries and unknowns. The whole point of science is to seek answers to those questions. Secondly, he tries to use arguments against evolution as evidence for intelligent design. Negative arguments against one theory do not necessarily provide evidence for a particular alternative. It would be like trying to show how safe a Honda Civic is by providing evidence that a Toyota Corolla is unsafe. No matter how unsafe the Corolla is, it says nothing about the Civic. Negative argumentation is scientifically useless without an alternative that can be positively tested. Despite these fallacies, it is still possible that Behe has raised an interesting and useful challenge to our current understanding of evolution.
In chapter 6 of Darwin’s Black Box, entitled "A Dangerous World", Behe outlines three IC biochemical systems in the immune system; clonal selection, V(D)J recombination, and the complement cascade. However, in his analysis of each of these systems, Behe makes critical mistakes. For the process of clonal selection, Behe targets the antibody molecules, which have the ability to rearrange their genes, and to switch from a membrane-bound form to a secreted form. Behe assumes that an antibody molecule missing one of these features would be useless. However, there are many examples of proteins that do not undergo rearrangement, but are able to switch forms. Additionally, there is a gene closely related to antibodies which can also undergo rearrangement, but does not switch forms. Behe neglects to mention either of these intermediate-like molecules. Instead, he scrutinizes the process of antibody gene rearrangement as the second IC system. He describes the numerous features of this system and claims there is no pathway through which it could have evolved. However, he assumes that the only selectable function for the genes involved in rearrangement is rearrangement. There are well-studied systems that bear remarkable similarity to V(D)J recombination that have nothing to do with immunity. He fails to recognize the existence of these other systems, nor realizes that most of the components of recombination are already present there. Finally, in his description of the third IC system, Behe mistakenly assigns a single function to the complement pathway. He fails to realize that the complement system has several different functions, and several mechanisms of activation. This would allow for a gradual, stepwise accumulation of parts and functions. In fact, many other groups of organisms possess a complement system that lacks his primary function. According to Behe’s definition of irreducible complexity, those systems shouldn’t exist.
In each case, Behe claims that a lack of articles in the scientific literature provides evidence that these systems could not have evolved. However, there are hundreds to thousands of articles published on these systems. Much was known about the evolution of each of these systems at the time Darwin’s Black Box was published. Additionally, much has been learned since then. This article will attempt to summarize the existing knowledge about the origins of these systems, and propose a possible pathway for their evolution through a Darwinian mechanism. It is not necessary to "prove" that these systems evolved. Because the essence of his argument is "how could this have evolved?", this article merely seeks to provide an answer to that question. One issue that will not be discussed is whether or not these systems are actually irreducibly complex. The term is not very useful to the discussion of origins because of the ambiguities surrounding its definition. In fact, IC as a term has failed to make much of an impression on the scientific community, as a search on the publication database PubMed, fails to turn up any references to "irreducible complexity"ii.
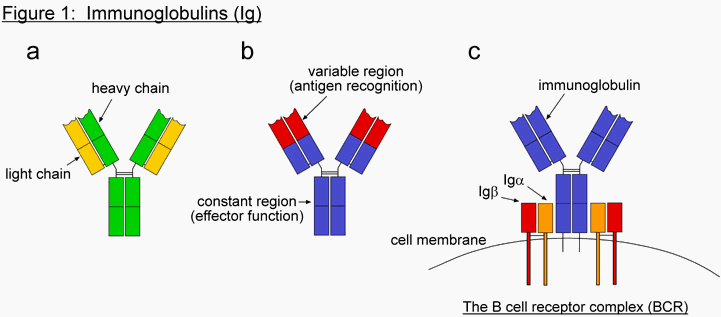
The first system Behe describes in chapter 6 is the process of clonal selection, which is how the adaptive immune system detects non-self molecules and initiates a response. This system focuses on a protein called immunoglobulin (Ig), and the complex it is part of, called the B cell receptor complex (BCR). Immunoglobulins, also called antibodies, are Y-shaped proteins that can recognize and bind foreign molecules (Figure 1a,b). They are produced by lymphocytes called B cells, and are initially expressed on the cell surface. In its membrane-bound form, immunoglobulins form a complex with two additional proteins, Ig
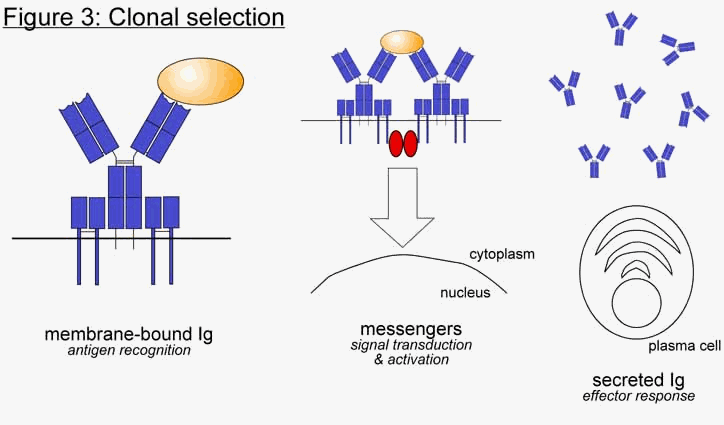
a and Igb, to form the BCR (Figure 1c). The targets of immunoglobulins are collectively referred to as antigen (Ag). Antigen can be essentially any molecule, though there is a preference for proteins. Because of the way in which immunoglobulins are produced, the specific antigen that each Ig recognizes is different in each B cell. When an unactivated, or naïve, B cell’s Ig recognizes its particular antigen, it sends a signal back into the interior of the cell to initiate activation. This signal is transmitted through a series of "messenger" proteins, which eventually enter the nucleus and transcribe a panel of genes involved in B cell activation. In order to become fully activated, further interaction with a second class of lymphocyte, called T cells, is required. Once the B cell is activated it proliferates and differentiates into plasma cells, and the function of the immunoglobulin changes from recognition to response. The immunoglobulin switches from a membrane-bound form (mIg) to a secreted form (sIg), where it can travel throughout the tissues of the host organism searching for pathogens. The only difference between the two forms is at the very tip of the bottom of the Y shape, which corresponds to the very tail end of the gene. Both forms are initially transcribed into mRNA, but through differential splicing only one form becomes translated into protein (Figure 2). The process of amplifying a particular B cell (or subset of B cells) from amongst billions based on its antigen specificity is called clonal selection (Figure 3). This process allows the immune system to choose from which B cell (or subset of B cells) it will mount an immune response. Additional discussion of the clonal selection system in the context of irreducible complexity can be found in the web article: The Evolution of Improved Fitness (http://www.talkorigins.org/faqs/fitness/)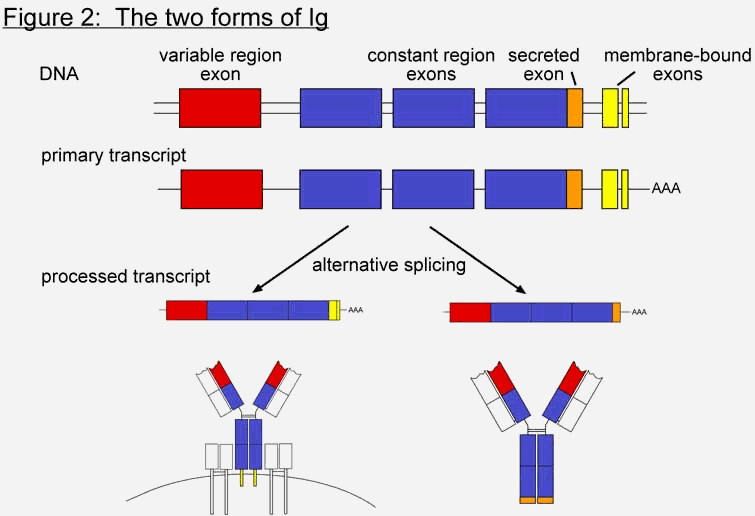
Behe summarizes this first system as a group of 3 components (page 125):
- membrane-bound form of the antibody
- messenger
- secreted form of the antibody
Behe has this to say about the evolution of the clonal selection system:
"A cell hopefully trying to evolve such a system in gradual Darwinian steps would be in a quandary. What should it do first? Secreting a little bit of antibody into the great outdoors is a waste of resources if there’s no way to tell if it’s doing any good. Ditto for making a membrane-bound antibody. And why make a messenger protein first if there is nobody to give it a message, and nobody to receive the message if it did get one? We are led inexorably to the conclusion that even this greatly simplified clonal selection could not have come about in gradual steps." (Page 125)
Switch forms or rearrange?
So the question is, could this have evolved? Certainly. One thing to realize is that the immunoglobulins are not the only antigen-receptors in the adaptive immune system. T cell receptors (TCRs) also undergo somatic gene rearrangement and can bind multiple and diverse antigens. However, the TCR only exists in a membrane-bound form, and is able to carry out its particular function without the need to leave the cell surface. Their receptors are used only for the detection of foreign antigens, and do not directly mediate an effector response. So even if immunoglobulins would be unable to effectively carry out their job if they were restricted to the membrane-bound form, they could have evolved from a TCR-like gene, which does not require secretion. If that occurred, then the ability to rearrange and the ability to switch forms could evolve in separate steps, with each step offering a selectable advantage.
It remains unclear whether a TCR-like gene appeared first or an Ig-like gene. However, it is also possible that the ability to rearrange came after the ability to switch. This would require that antigen receptors with a single specificity be effective mediators of immunity. Could this have occurred? Absolutely. There are several families of innate receptors that do not rearrange, called pattern recognition receptors (PRRs) (Medzhitov and Janeway 1997). Bacteria possess many unique and essential surface molecules that are distinct from metazoan surface molecules. For example, organisms ranging from insects to humans bear numerous receptors capable of recognizing the bacterial cell wall component lipopolysaccharide (LPS) (Fenton and Golenbock 1998). Examples include the macrophage scavenger receptor, CD14, and the
b2-integrins. Binding to these receptors induces a variety of responses, from the release of inflammatory agents to the phagocytosis of the LPS-carrying particle. Some of these receptors are membrane-bound and some are secreted. Interestingly, there are some that can exist in both forms, such as CD14. So here we have examples of receptors that do not rearrange, can be membrane-bound, secreted, or both, and can mount an effective immune response.Recently, a protein was discovered in the silkmoth Hyalophora cecropia that is upregulated upon exposure to LPS (Sun, Lindstrom et al. 1990). This protein, called hemolin, is secreted into the hemolymph (the primitive invertebrate "blood") and can bind to the most toxic region of LPS (Daffre and Faye 1997). This protein is also constituitively expressed in a membrane-bound form on the surface of hemocytes. Interestingly, the structure of this protein was found to contain 4 Ig domains, and is a member of the Ig superfamily (Lindstrom-Dinnetz, Sun et al. 1995). Ig domains are a type of 3D protein structure first identified in immunoglobulins, consisting of 7 or 8
b-pleated sheets. Additionally, the crystal structure was recently solved and bore a shape loosely similar to one of the arms of immunoglobulins (Su, Gastinel et al. 1998). However, sequence analysis showed that hemolin most likely does not represent a precursor to immunoglobulins as the type of Ig-domain it possessed (the I-set) was one not found in vertebrates. Hemolin does show similarities both structurally and functionally to cell-adhesion molecules, and also contains some cell-adhesion activity. This dual function, both for immune response and cell-adhesion, suggests that immune molecules arose from cell-adhesion molecule precursors. Hemolin is a clear example of an inducible immune molecule that can exist in both membrane-bound and secreted forms.
Immunoglobulin signal transduction
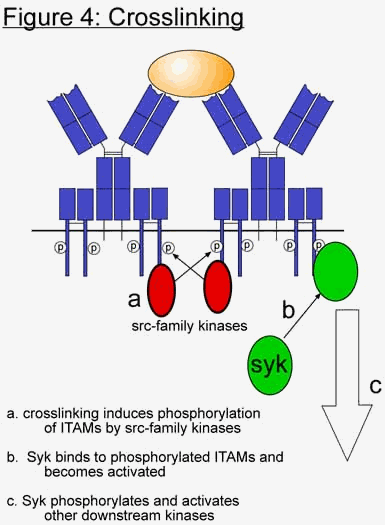
When an antibody recognizes an antigen, it sends a signal back into the cell, which "primes" the cell for activation. But what is the nature of this signal? How exactly does the binding of antigen, which occurs on the exterior of the cell, induce a signaling cascade on the interior? The answer lies in the branched nature of the immunoglobulin. As previously stated, immunoglobulins are loosely shaped like a Y, with two arms sticking out into the extracellular space, and the bottom embedded into the membrane. Associated with each Ig are the two accessory proteins Ig
a and Igb. These accessory molecules have long cytoplasmic tails, and contain motifs specialized for attracting the type of kinases required to initiate the cascade. These motifs are called ITAMs, which stand for immunoreceptor tyrosine-based activation motifs. These motifs serve as targets for protein tyrosine kinases, which phosphorylate the tyrosine residues on the ITAMs. Because most antigens have multiple epitopes (i.e. multiple binding sites for antibodies), and because the immunoglobulins have multiple arms, the recognition of an antigen results in a clustering of immunoglobulins on the membrane in the area where the antigen is located (Fig 4). This process is often called crosslinking, and is the key to activating the cell. Visually, it looks kind of like a "hands across America", where each arm of the Ig is linked to an arm of another Ig, and so on and so on. As more and more Igs cluster around the antigen, a family of kinases, called the src family kinases, which are loosely associated with the cytoplasmic tails of Iga and Igb, begin to phosphorylate the ITAMs. Because the src kinases are attracted to phosphorylated tyrosines, this sets up a positive feedback loop, and more and more src kinases are recruited to the site. In addition to attracting themselves, they also attract another kinase called syk. The newly bound syk kinases begin to phosphorylate and activate other syks, and propagate the cascade through other signaling molecules. These other signaling molecules are expressed in many different tissues and are considered to be more generic transducers of signaling cascades. This leads to the activation of transcription factors (TFs), which then enter the nucleus and initiate the transcription of genes involved in cell proliferation and differentiation. In this particular situation, the B cell becomes "primed" for activation by T cells.Most of the "messengers", or components of the signaling pathway are homologous to many other signaling pathways present in other tissues and in other organisms. For instance, one of the final components in the pathway is the activation of the transcription factor NF-
kB. This transcription factor is used in multiple tissues in a wide variety of organisms to transcribe numerous immune-related genes. The recognition sequence for NF-kB is GGGRNNYYCC (where R = purine, Y = pyrimidine, and N = any base). Interestingly, it is also a key TF in the insect immune system, and is activated by the toll receptor pathway leading to transcriptional activation of many insect immune-related genes. Even in mammals many innate receptors, including the LPS receptor CD14, activates transcription through NF-kB. This also proceeds through a toll-like receptor. For an antigen-receptor to co-opt a pathway such as this, it would only need to link its antigen-receptor to the start of that network. This could be achieved by acquiring accessory molecules with ITAMs. However, until recently it was thought that ITAMs only existed in accessory molecules to Igs and TCRs. But recently, a protein that contains two ITAMs was discovered in the invertebrate ascidians (tunicates) (Takahashi, Ishikawa et al. 1997). This protein is required for cellular defense, such as phagocytosis, as is mediated by hemocytes (tunicate "blood" cells). Additionally, more recent experiments demonstrated that the activity of this protein is dependent upon tyrosine phosphorylation (Ishikawa, Azumi et al. 2000). While the tyrosine kinase responsible for phosphorylating the ITAMs of this protein remains to be discovered, it strongly suggests that a precursor system, with ITAM-bearing accessory molecules, tyrosine kinases, and NF-kB, was already in place long before the immunoglobulins appeared.
Using the information presented, a scenario for the evolution of the immunoglobulin detection/secretion pathway can be postulated. (1)It begins first with an innate membrane-bound receptor that can induce the transcription and release of antimicrobial peptides upon interaction with a specific foreign antigen (such as LPS). (2)It then evolves the ability to switch to a secreted form to directly inhibit the antigen-bearing pathogen. (3)Finally, it evolves the ability to rearrange itself to produce a diverse array of specificities. This scenario is fully consistent with the known literature, and the provided examples demonstrate the feasibility and selectability of each step: (1)The LPS receptor in drosophila (or mammals), (2)hemolin in moths, and (3)immunoglobulins.
IC system II: The Rearranging Antigen Receptor Loci
There are on the order of one billion B cells circulating in a human being at any given time. Each of these B cells expresses an immunoglobulin with a different specifity (actually, the surface of the B cell has several thousand immunoglobulin proteins attached to it). How is this diversity acheived? The genes encoding immunoglobulins (and TCRs) are composed of multiple gene segments, which are cut-and-pasted together in each B cell to form a functional Ig gene through a process called V(D)J recombination, or gene rearrangement (Figure 5). In non-lymphocytes, which do not undergo rearrangement, the gene segments are clustered in 2 or 3 groups. For example, at the Ig heavy chain locus, there is a cluster of about 50 V gene segments, a second cluster of 27 D gene segments, and third cluster of 6 J segments. During B cell development, the cell will randomly choose one gene segment from each cluster, and attach them together to form a continuous V-D-J gene. This forms the antigen-binding region of the heavy chain. From this process alone, 8100 different combinations of heavy chains can be produced. In addition, the joining process is imprecise, and several nucleotides can be added or removed at each junction. These junction points actually form a portion of the antigen-binding pocket at the tip of the immunoglobulin.
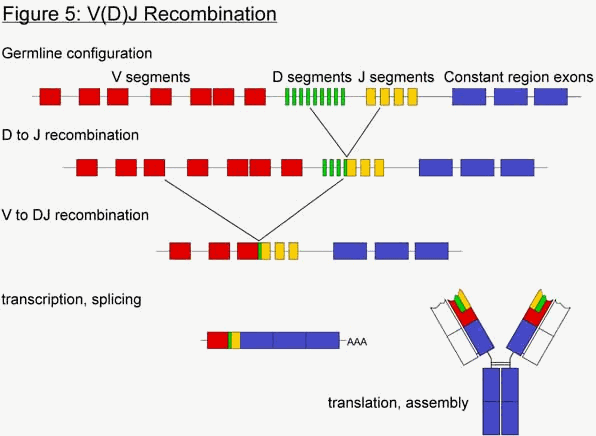
In the last 12 years, numerous experiments have shed great light on the mechanism of V(D)J recombination. It essentially involves two proteins, called RAG-1 and RAG-2. RAG stands for Recombination Activating Gene. These two proteins together recognize special sequences flanking each gene segment, called recombination signal sequences (RSS). The RAG proteins bind to the RSSs of two gene segments, then bring them in close proximity to each other and cleave the DNA precisely at the junction between the gene segment and the RSS (figure 6). DNA repair complexes then repair the double-stranded breaks, but attach the two gene segments to each other, and the two RSSs to each other. The region in between the gene segments, which include the RSSs, ends up getting deleted in the process. The DNA of the B cell is therefore irreversibly altered by the recombination event.
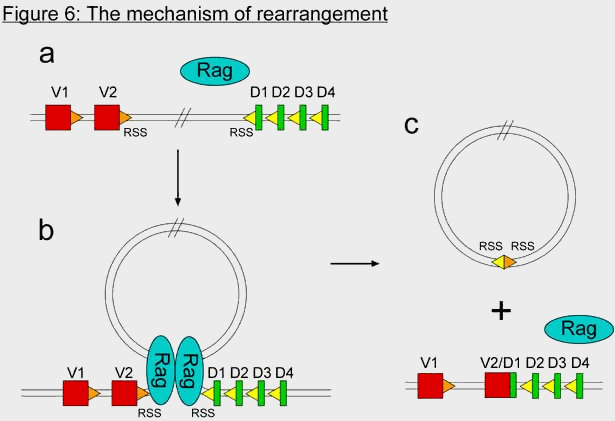
According to Behe, an antibody-diversity system requires three components: the antibody genes themselves, the recombination signal sequences (RSS), and the recombination activating genes RAG-1 and RAG-2.
"In the absense of the [RAG] machine, the parts never get cut out and joined. In the absense of the signals, it's like expecting a machine that's randomly cutting paper to make a paper doll. And, of course in the absence of the message for the antibody itself, the other components would be pointless." (Page 130).
Behe paints a pretty dire picture for how a process like this could possibly have originated in Darwinian fashion. He even goes further to state:
"The problem of diversity runs headlong into the requirement for minimal function. A primitive system with only one or a few antibody molecules would be like the propeller turning at one revolution per day: not sufficient to make a difference... Because the likelihood is so small for the shape of one antibody being complementary to the shape of a threatening bacterium -- perhaps one in a hundred thousand or so -- any animal that spent energy making five or ten antibody genes would be wasting resources..." (Page 130-1).
What Behe neglects to mention is that numerous non-rearranging, innate receptors already exist in the immune system. The LPS receptor is only one of many non-rearranging receptors that play an important role in immunity. Others include the glycan receptor, the mannose receptor, and the macrophage scavenger receptor; all of which recognize many different bacterial constituents. The activation of an adaptive immune response begins about 3 days after the start of infection. Until those three days have passed, the only thing keeping the pathogens at bay is the innate immune system. Phagocytic cells like macrophages and neutrophils express a wide array of innate receptors that allow them to detect and engulf many types of pathogens before they have a chance to multiply and overwhelm the immune system. So Behe is quite wrong to conclude that non-rearranging antigen receptors are not beneficial to the host. Invertebrates, which lack an adaptive immune response, are entirely dependent on innate receptors for survival. What is key to remember is that bacteria are very different from their metazoan targets. There are numerous surface molecules essential to bacteria that are absent in animal cells. These differences have been capitalized upon by the innate immune system hundreds of millions of years prior to the emergence of the adaptive immune system.
Assuming that a non-rearranging antibody-like gene is beneficial, there still leaves two important components that needed to appear during the evolution of antibody diversity; the signal sequences and the RAG genes. Their origin is indeed a mystery. However, many clues can be discovered merely by comparing the mechanism of recombination to other biochemical processes. The process of recombination itself bears a great deal of non-trivial similarity to another process which has nothing to do with immunity. In fact, this process could be considered to be on the opposite end of host protection. It is a process utilized by bacteria for antibiotic resistance, and by retroviruses to invade the genome of their hosts. This process is transposition.
A transposable genetic element is a region of DNA capable of "jumping" from one location on the genome to another. These regions are around the size of a gene, as they usually encode the protein that catalyzes the transposition reaction. Flanking these regions are sequences that facilitate transposition, usually consisting of inverted repeats. While the precise mechanism of transposition varies from system to system, it typically involves the nicking or cleavage of DNA at sites on both the target and donor, then swapping the two strands to integrate the transposon into the target DNA. This process can be conservative or replicative, meaning that it may or may not leave a copy of the transposon at its former location. The exact function of transposons remains unclear, but they exist in numerous organisms, including bacteria, viruses, corn, worms, and fruit flies. Their presence in such a wide variety of lifeforms suggests that either they are functionally important, or extremely efficient parasites. A role for mobile elements has even been implicated in evolution (Hall, Collis et al. 1999).
V(D)J recombination shares a lot of similarity to transposition (Lewis and Wu 1997; Lewis 1999; Schatz 1999). The RAG genes are tightly clustered on the genome, being only 8 kb apart (Oettinger, Schatz et al. 1990). Additionally, each gene is encoded by a single exon. This type of genomic organization is rare for higher eukaryotes, but common for genes contained within transposons. Additionally, the RSS sequences are similar to the inverted repeat sequences used by many transposons (Dreyfus 1992). The reaction itself has many features similar to transposition (Spanopoulou, Zaitseva et al. 1996). For instance, the cleavage reaction mediated by the RAGs leads to the formation of a hairpin intermediate structure (Roth, Menetski et al. 1992). Transposases, such as the Tn10 transposase in bacteria, can also generate hairpin intermediates (Kennedy, Guhathakurta et al. 1998). This feature is also shared by retroviruses during integration (Van Gent, Mizuuchi et al. 1996). However, the strongest piece of evidence favoring a transposase origin for V(D)J recombination was the demonstration of transposase activity by RAGs in vitro (Agrawal, Eastman et al. 1998; Hiom, Melek et al. 1998). With that discovery, there could be no doubt that the similarities between the two processes wasn’t merely circumstantial.
How can we use this information to generate a model for the origin of V(D)J recombination? Behe states:
"As scientists we yearn to understand how this magnificent mechanism came to be, but the complexity of the system dooms all Darwinian explanations to frustruation." (Page 139).
It’s a good thing not all scientists are as easily discouraged as Behe by the complexity of this system. Let’s see what others have come up with. The transposon model is currently the most feasible model to explain the origin of V(D)J recombination. The model essentially states that the RAGs were originally contained within a transposon, and that this transposon integrated into a primordial antigen receptor (Thompson 1995). Later the RAGs were moved to another location on the genome, where their expression could be better regulated. The initial integration event, in one step, inserted both the RAG genes and the RSS sequences, and generated a rearranging antigen receptor.
A more detailed model could proceed as follows: A transposon containing the RAG genes and flanked by RSSs integrates itself into the primordial antigen-receptor gene, splitting it into two gene segments (V and J). The locus itself is silenced in most cells, preventing expression of the RAG genes and removal of the integrated transposon. However, in a lymphocyte-like cell, the locus becomes transcriptionally active, and the RAG genes express themselves and remove the transposon, reuniting the two gene segments. The imprecise joining process generates a level of receptor diversity that favors the organism bearing this transposon, and its descendants thrive with the increased immune capabilities. A gene duplication creates a second locus, and the two lineages become the heavy and light chains. A second integration event by another transposon splits the heavy chain’s V segment into V and D segments. Further gene duplications create additional V, D, and J gene segments, giving the receptors multiple rearrangement targets and increasing their diversity of antigen recognition.
The similarity between RAGs and transposases is not a recent discovery. Similarities between the two systems have been known as early as 1979, when the RSSs of the kappa light chain J segments were first sequenced (Sakano, Huppi et al. 1979). In the discussion section, the author stated, "We propose that such a mechanism was initiated when an IS-like DNA element was accidentally inserted into one of the multiple V DNA copies of an ancestral polymeric gene,.." While the model has been greatly refined since then, the basic concept remains the same. Why would Behe neglect to mention the transposon-origin model? Sakano et al is cited by 706 other research articles, clearly it is taken seriously by the scientific community. Instead, Behe cites a two-page commentary in PNAS (Bartl, Baltimore et al. 1994) as the "best" article discussing the molecular origins of V(D)J recombination, to which he describes as making "a valiant stab at accounting for the components, but in the end, it is a hop in the box with Calvin and Hobbes." He also adds, "their proffered solutions are really just a disguised shrug of their shoulders." He makes no attempt to address their model, dismissing it with a wave of his hand. He also fails to appreciate the context of the article, which is primarily a commentary on how the discovery of TCR genes in sharks two months earlier had influenced current opinion. They did not delve into the details of the transposon-origin model not only because the scope of the article was so focused, but also because so many articles before had described it. Their aim was to add to the discussion, not reiterate it. They assumed the reader would already be familiar with the basics of that model. Rather than stopping after reading one article, perhaps Behe should have used the "related articles" tool on PubMed and read one of the other 220 articles listed. For example, the article "New Insights into V(D)J Recombination and It’s Role in the Evolution of the Immune System", is cited by 90 other articles (Thompson 1995). This 9-page review article provides a much more in depth description of the transposon-origin model. Behe was quite incorrect when he stated:
"We can look high or we can look low, in books or in journals, but the result is the same. The scientific literature has no answers to the question of the origin of the immune system." (Page 138)
This proclamation is absurd, and academically irresponsible. Behe singles out a commentary for scrutiny, ignoring the wealth of literature on the topic, and concludes no answers exist. He should have looked harder, or asked an immunologist. Hypotheses put forth at the time of Darwin’s Black Box’s publication have since been confirmed, and the transposon-origin model is now quite robust, especially with the discovery of the transposase activity in RAGs. If only ID could claim similar progress in the last 6 years.
The complement system is a cascade of serine proteases initiated by the detection of foreign agents, and resulting in several different effector responses. A serine protease cascade is system of proteins that can sequentially activate each other through cleavage, and possess a serine residue in their catalytic site. They function analogously to a chain letter, where the first protease cleaves and activates several target proteases, who in turn cleave several of their targets, and so forth. The result is an exponential amplification of activity, set off by as few as a single activated protease. In mammals, the cascade as a whole uses around 30 different proteins, which are all constituitively expressed in the blood serum. In fact, complement proteins are believed to comprise a higher percentage of blood plasma proteins than any other group of proteins.
There are essentially three ways to initiate the cascade, called the classical, alternative, and lectin pathways (Figure 7, top boxes). While the classical requires the binding of antibody to antigen, the alternative and lectin pathways do not. The classical pathway was the first of the three to be discovered, due to its ability to "complement" the activity of antibody molecules in eliminating red blood cells in cross type blood transfusions. When multiple antibody molecules recognize its target, the complement factor C1 (which is a complex of three different proteins, C1q, C1r, and C1s), binds to the antibody and becomes activated. It can then catalyze the cleavage of C4 into C4a and C4b. C4b contains a highly reactive thioester group, which causes it to covalently bind to proteins near it, which presumably includes the antibody’s bound target, like the surface of a bacterium. C2 can then attach to C4b, and is then cleaved by C1 into C2a and C2b. C2b and C4b then form the second serine protease in the cascade, called C3 convertase. This complex induces the cleavage of another factor, C3, into C3a and C3b. C3b can group with the C4b/C2b complex to form a C5 convertase complex, which cleaves C5 into C5a and C5b. C5b attracts the complement proteins C6, C7, C8 and C9. This group of proteins, C5b through C9, forms a membrane attack complex, which is basically a pore on the surface of the bacterium. The formation of several of these pores disrupts the osmotic balance of the bacterium, resulting in its lysis.
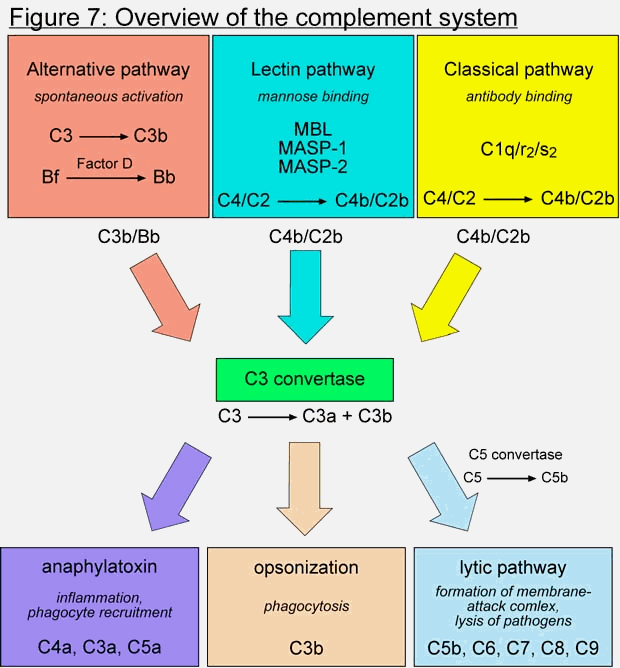
Aside from his typical complaint that this system is "too complex to have evolved", Behe does little to actually specify what makes the complement pathway irreducibly complex, and does not give a list of essential components. He spends a lot of time describing the classical pathway, and its requirement for antibody-binding. By focusing on the classical pathway, Behe implies that antibody-binding is essential to complement activation. This could not be further from the truth. Organisms lacking an adaptive immune response, such as lampreys or tunicates, do posses complement factors and have an efficient complement system (Nonaka and Takahashi 1992; Smith, Azumi et al. 1999). These animals do not use the classical pathway of activation, but rely instead on the lectin and alternative pathways. In mammals, the lectin pathway is initiated by the binding of a protein called MBP (mannose-binding protein) to a mannose-containing saccharide on the surface of many types of gram-negative bacteria (Matsushita, Endo et al. 1998). This leads to the binding of two proteins, MASP-1 and MASP-2 (MBP associated serine protease), which catalyze the cleavage of C4 and C2 into C4a/C4b and C2a/C2b, which then joins the classical pathway at the C3 convertase. Lampreys and tunicates have similar lectin pathways as mammals. Interestingly, the complex of MBP/MASP-1/MASP-2 very strongly resembles the C1q/C1r/C1s complex, in sequence, structure, and function.
The alternative pathway is much more simple to activate, utilizing the fact that an intermediate step in the classical and lectin pathways, the cleavage of C3, can often occur spontaneously. As it turns out, C3b can associate with a protein called Factor B (Bf) to form a C3 convertase, eliminating the need for C4b and C2b entirely (Xu, Narayana et al. 2001). Factor B binds to C3b, and is then cleaved into its active form, Bb, by a serine protease called Factor D. The mechanism for activation is quite similar to C4b and C2b, and C3b is very similar in structure to C4b, and Bb is very similar to C2b. Additionally, Factor D has sequence similarity to trypsin, a very common serine protease. Behe leaves out the lectin pathway entirely, and only briefly mentions the alternative. Why? It is clear that the origins of the complement system lie within one of these two activation pathways, so their function and history should be focused upon. By spending so much time with the classical pathway, Behe makes the system’s origin appear more complicated than it actually may be.
Behe defines an irreducibly complex system as a system that contains multiple essential parts, where the elimination of any one of these parts results in the loss of the function of that system. However, in the case of the complement system, their function is somewhat unclear. Behe never states specifically what the function of this pathway is, but implies that it is to lead to the formation of the membrane attack complex (MAC), thereby causing the lysis of the bacterium. However, the branch of the cascade leading to MAC formation (called the lytic pathway) represents only one particular effector function of complement. In general, the complement pathway leads to three types of effector responses; inflammation, opsonization, and MAC formation (Figure 7, bottom boxes). While the cleavage of C2, C4, C3, and C5 is important for their function in propagating the cascade, the cleavage products C4a, C3a, and C5a are all anaphylatoxins. These function as local mediators of inflammation, which can recruit innate immune cells like phagocytes to the site of infection. C2b, C4b, C3b, and C5b can also serve as opsonins. That is, they can facilitate the phagocytosis of their bound targets by binding to receptors on the surface of phagocytes. Since the lytic pathway has so far not been detected in organisms lacking an adaptive immune system, the opsonizing function of complement is presumably its original function. However, Behe doesn’t even mention the complement system’s function in opsonization. Nor does he recognize the fact that many organisms that lack both the classical pathway of activation and the lytic pathway still have a robust complement-mediated immune system. It would seem that his intention here is to snowball the reader with endless descriptions of complexity, rather than specifying the key features that directly relate to evolution and origins. Additional discussion of the complement system as an irreducibly complex system can be found in the talkorigins article, "
Is the Complement System Irreducibly Complex?"
"Comparing sequences might be a good way to study relatedness, but the results can’t tell us anything about the mechanism that first produced the systems." (Darwin’s Black Box, Page 138).
Sequence comparison is one of the first steps in analyzing the function of a gene. By comparing the sequences of a gene between two distantly related groups, regions important for function can be identified for future experiments. This is a commonly used strategy in any lab. Comparing the sequences of a family of genes within multiple groups of organisms can reveal a wealth of information on the evolution of that family, including which members appeared through gene duplications and when. All of this information can be used to infer the function of the ancestral state of the system, and how it might have originated. This is a much more effective means of examining the origin of a system than Behe’s apparent method, removing components and testing for loss of function. Removing components is not like rewinding a tape. It does not necessarily tell us what the ancestral state was like.
While a fish probably looks more like the common ancestor of mammals and fish than we do, it is still over 100 million years more evolved than its ancestor. However, we can still look at the features of a fish’s immune system and compare it to our own. If a similar feature is observed, it most likely arose prior to the divergence of the two groups. By comparing humans to other groups, we can begin to trace back our evolutionary roots. The following section attempts to explore the innate immune systems of other organisms, in relation to our own. It begins with more distantly related organisms and continues through to the jawed vertebrates. Though separated by 450 million years of evolution, even cartilaginous fish like sharks and manta rays have essentially the same complement pathway as humans (Nonaka and Smith 2000). Refer to figure 8 to see the relationships between each group.


Arthropods (phylum Arthropoda)
Insect immunity is mediated primarily through the secretion of antimicrobial peptides, whose exact functions are unknown. While insects do not have a complement system, they do possess several different types of lectins and serine proteases, which have features similar to complement and blood-clotting. Some lectins are able to induce coagulation upon recognition of certain microbial surface constituents (e.g. LPS) (Muta and Iwanaga 1996). Coagulation is a serine protease-dependent process that forms an insoluble matrix surrounding infectious particles, which are then cleared from the hemolymph (insect "blood"). This process has many features in common with blood-clotting. Furthermore, lectins with fibrinogen-like domains, such as Tachylectin 5A and 5B (TLs-5) are also involved in this process (Gokudan, Muta et al. 1999). Fibrinogen is one of the key players in vertebrate blood-clotting. TLs-5 may represent an evolutionary link between immunity and blood-clotting (Kairies, Beisel et al. 2001).
a
2-macroglobulin (a2M), a protease inhibitor, can regulate the hemolytic activity of proteolytic enzymes in the hemolymph (insect "blood"), such as the lectin limulin (Armstrong and Quigley 1999; Swarnakar, Asokan et al. 2000). It contains a protease sensitive region, which serves as "bait" for a wide variety of proteases. Once cleaved, a highly reactive thioester bond is exposed, which covalently attaches a2M to the protease. a2M is then able to form a "cage" around the protease, which is then cleared from the hemolymph through receptor-mediated endocytosis (Melchior, Quigley et al. 1995). As such, a2M can not only regulate its own proteases, but also deactivate pathogenic proteases secreted by infectious agents. a2-macroglobulin shows remarkable homology to C3, C4, and C5, suggesting that a2M may represent an evolutionary precursor to C3/C4/C5 (Sottrup-Jensen, Stepanik et al. 1985). Remarkably, a thioester protein similar to a2-macroglobulin was discovered recently in mosquitos, called aTEP-I (Levashina, Moita et al. 2000). aTEP-I has complement-like opsonization activity, and could represent an evolutionary precursor to C3. Furthermore, it had weak binding activity in the absence of proteolytic activation, suggesting that a more ancestral form of this protein could function without being cleaved.Other immune-related molecules include the pentraxin lectin family. One of these components, C-reactive protein (CRP), is present in vertebrates, and there can activate the classical complement pathway (Armstrong, Armstrong et al. 1993). Overall, it appears that the very beginnings of complement-mediated immunity as well as blood-clotting may have begun before the proteostome/deuterostome split.

Echinoderms (phylum Echinodermata)

Urochordates (phylum chordata, subphylum urochordata)
Ficolins are proteins composed of basically two domains, a fibrinogen-like domain used for carbohydrate binding, and a collagen-like domain used to activate the MASPs (Matsushita and Fujita 2001). MBP also possesses a collagen-like domain very similar to ficolins, but does not have a fibrinogen-like domain. In addition to activating complement, ficolins and MBP can act as opsonins through their collagen-like domains. This suggests that ficolin or MBP may have been added to the already existing alternative pathway to form the lectin pathway in an ancestor to the urochordates.
Cephalochordates (phylum chordata, subphylum cephalochordata)
Amphioxus (also called lancelets) of the subphylum cephalochordata are considered to be the closest extant relative to the vertebrates. Their basic shape and genomic organization resemble vertebrates, but they are much simpler. As a result, they are often studied to shed light on key features of vertebrate development. Although immunological research on amphioxus is just beginning, a key discovery has already been made. A C3-like gene and a C6-like gene were found in the amphioxus, Branchiostoma belcheri (Suzuki, Satoh et al. 2002). While the C3-like gene is not surprising, the presence of a C6-like gene is. C6 is a component of the cytolytic membrane-attack complex (MAC), which is involved in the lysis of foreign pathogens. This strongly suggests the presence of a lytic pathway in cephalochordata. The complement proteins C6, C7, C8, and C9 all share key structural motifs, which may indicate a common origin. Phylogenetic comparisons of amphioxus C6 to human C6, C7, C8, and C9 suggest that amphioxus C6 is not that common ancestor. These results would therefore predict the existence of additional MAC components in amphioxus, and that the lytic pathway has a more ancient evolutionary origin

The jawless fish, or agnathans, of the class cyclostoma has only two extant groups, lampreys and hagfish. They represent the most ancestral vertebrates. Despite extensive searching, no rearranging antigen receptors or MHC genes have been found in cyclostomes. On the basis of this negative evidence it is generally assumed that they do not possess an adaptive immune system, making them the only group of vertebrates without one. However, and a C3 homologue has been detected in lamprey, as well as a Bf/C2 homologue, and a MASP homologue (Nonaka, Fujii et al. 1984; Nonaka, Takahashi et al. 1994; Endo, Takahashi et al. 1998). The lamprey Bf/C2 gene is equally related to mouse Bf and mouse C2, indicating that the gene duplication that produced mouse Bf and C2 occurred after the cyclostomes diverged from the jawed vertebrates. Lamprey C3 is more similar to mouse C3 than to mouse C4 and C5. However, a lamprey C4/C5 homologue has not yet been identified, so it remains unclear whether the cyclostomes emerged before or after the C3/C4/C5 gene duplications (Nonaka 2001). Interestingly, lamprey C3a is very similar to mouse C3a, while ascidian and sea urchin C3a are not. Since mouse C3a serves as an anaphylatoxin (an inflammatory mediator), it suggests that this ability evolved in an ancestor to the cyclostomes. Additionally, a C5a-like anaphylatoxin ability has been reported in hagfish (Newton, Raftos et al. 1994). The lytic pathway of the complement system has not yet been observed in cyclostomes; however, a gene was cloned recently in hagfish that bears remarkable similarity to CD59 (dos Remedios, Ramsland et al. 1999). In mammals, CD59 is an inhibitor of the lytic pathway, preventing lysis of self cells. The cloned gene has 33% amino acid similarity to CD59, suggesting that it is indeed a homologue. Recent data also suggests that lampreys have lymphocytes, but since lymphocytes in all other vertebrates utilize rearranging antigen receptors, their role in lamprey immunity remains unclear (Shintani, Terzic et al. 2000).
In summary, it appears that the jawless fish contain all the similar immune related genes and pathways that sea urchins and urochordates possess. However, several new features have appeared, such as the use of anaphylatoxins, the possible appearance of lymphocytes, and preliminary evidence that suggests the existence of the lytic pathway. Unfortunately the next group of organisms, from the class Placodermi, are all extinct. Unless we develop the technology to extract DNA from their remains, their immune systems will most likely remain a mystery.

Cartilaginous fish (phylum Chordata, class Chondrichthyes)
The cartilaginous fish generally includes three groups of organisms; sharks, skates, and rays. They are classified due to their cartilaginous skeletons. They also possess jaws, and are the most ancestral group of the jawed vertebrates, or gnathostomes. They are also the most ancestral organisms that possess an adaptive immune system. Their innate immune systems possess all three complement activation pathways (classical, alternative, lectin), and their complement can serve as anaphylatoxins, mediate opsonization, and activate the lytic pathway (Smith 1998). While not all components homologous to the mammalian complement system have yet been identified, it is clear that their system is as complete as ours. It is likely that in the coming years all components will be discovered. In that sense, the story of the origins of complement-mediated immunity ends with the cartilaginous fish.
Any model for the origin of the complement system must begin with the evolution of an
a2-macroglobulin-like gene into a C3-like gene. C3 is the key molecule in the complement system. It can interact with as many as 25 other proteins, and is the point of junction for all three activation pathways (Sahu and Lambris 2001). C3’s strong sequence similarity to a2M, which is not part of the complement system and presumably more ancient, clearly suggests a possible origin. Additionally, C3 is not a serine protease, and an explanation is required for how it became associated with serine proteases. The fact that a2M is a protease inhibitor and serves as a target for bacterial proteases also suggests how this may have come about.In the scientific literature, the models that are proposed for the evolution of the complement system are not nearly as detailed as the following model. The reason for that is simple. Behe is the only person that is requiring that level of detail in order to be convinced that it could have evolved. It would be similar to hypothesizing that a particular football team would win the superbowl, but then a critic would demand details like how many first downs the team would get, the total yardage accumulated by both offenses, whether Gatorade would be poured upon the coach, etc. That level of detail is not justified based on what information is currently available. Therefore, the following model is not meant to be completely accurate. It’s an attempt to synthesize the current information to provide an example of what could have happened, not what did happen. According to Behe, a Darwinian explanation for the evolution of an irreducibly complex system is a logical impossibility. Therefore, to falsify his IC to ID inference, we need not prove the model. We need only to provide a reasonably plausible model.
It should also be mentioned that this model contains many holes, many gaps that need to be filled by experiments or other evidence. However, the model also makes predictions, some of which can lead to future research. Future experiments may disprove this model. That is perfectly fine, it will just allow us to generate a new model that will fit the new evidence. This new model will make further, more specific predictions, and the cycle will continue until all remaining holes have been filled. We’re at the beginning of the genomic era, and the comparison of sequences will allow us to test and refine this model in ways that would have previously been impossible.
The origin and evolution of the complement system
a2-macroglobulin. It was constituitively expressed in the hemolymph of the early bilaterians around one billion years ago. The key to its function was the protease-sensitive "bait" region, which when cleaved exposed a highly reactive thioester site, which would covalently bind to whatever protease cleaved it. After attachement, the a2M/protease complex would be cleared from the system through phagocytosis. This would provide some protection for the host from various types of bacterial proteases.The evolution of complement begins with an ancestor to protease inhibitor
Numerous carbohydrate-binding lectins are present in the hemolymph of all bilaterians. Some of these can recognize bacterial cell wall constituents (e.g. LPS) and serve as opsonins. Shortly after the deuterostomes split off from the protostomes, a serine protease domain was attached to a carbohydrate-binding domain through exon shuffling. This protein, a precursor to ficolins, could bind to bacteria, then cleave and activate the
a2M-like protein. Because a2M binds covalently to its target, this would give a much stronger opsonizing activity than through lectin binding alone. A gene duplication of a2M and subsequent differentiation through random mutation followed by natural selection would optimize the interaction between the ficolin precursor and the protein that would eventually become C3. Because of the reactivity and lack of specificity of the C3 precursor, various inhibitors of C3 began to evolve on the surface of host cells, preventing them from being engulfed.The key step in creating the complement pathway began sometime before the echinoderms branched off from the prechordates. The ancestral C3 protein evolved an affinity for a serine protease, which would attach to C3b after cleavage. By localizing a serine protease near the site of C3b attachment, the host could generate more C3b in close proximity to the pathogen. With the addition of the ancestral Factor B, the positive feedback loop that would be the hallmark of the complement pathway was born. This loop gave the ancestral complement system the ability to deposit large amounts of C3b onto the surface of pathogenic molecules very rapidly. Additional evolution produced greater specificity through an increased affinity for C3b by itself, and better regulation by the requirement for C3b-binding for Factor B activation.
Because of the numerous regulatory molecules preventing complement activation against self-molecules, the requirement for lectin activation of complement became increasingly less stringent. C3 could evolve to become more and more spontaneously reactive, as low levels of serum C3b were tolerated. With the addition of a dedicated activator in Factor D, the alternative pathway was created.
After the echinoderms branched off from the chordates, more sophisticated lectin complexes evolved. The MASPs were created, possibly from a gene duplication of Bf (Nonaka and Miyazawa 2002). They originally interacted with ficolins, but through an exon shuffling event joining a ficolin’s collagen-like domain to a CRD, mannose-binding protein (MBP) was created. Sometime before the emergence of the jawed vertebrates, a genome-wide duplication event created additional copies of C3 and Bf. These would later evolve into C4 and C2. The lectin pathway would then utilize these new genes, increasing their affinity for C4 and C2, and decreasing their affinity for C3 and Bf. By adding an additional step, the amplification power of the protease cascade was increased.
When the adaptive immune system appeared with the jawed vertebrates, the complement system was quick to take advantage of it. Gene duplications of MBP and the MASPs created C1q, C1r, and C1s. Once C1q evolved the ability to bind to immunoglobulins instead of carbohydrates, the joining of these two pathways was complete. The combined system possessed the specificity of immunoglobulins, and the effector response of complement.
The addition of the lytic pathway to the complement system could have occurred before or after the creation of C4. One of the major features of any model of complement evolution is the notion that C4 and C5 were created from gene duplications of C3. However, it remains unclear whether C4 was created first, or C5. In jawless vertebrates, only C3 is present. In jawed vertebrates, both are present. Regardless, at some point either before or after the emergence of the jawed vertebrates, the lytic pathway was added to the complement system. C5 was initially created, and had opsonizing activity (C5b), and anaphylatoxin activity (C5a). A molecule with a perforin domain, which is the key domain in mediating cell lysis, appeared. In time, this perforin-like gene evolved into C6, C7, and C8, and could lyze enucleated cells. C9 was added as a final step to form a pore capable of lysing nucleated cells.
What predictions can be made from this model? One interesting feature of this model is that it suggests that the ancestral complement system required a lectin pathway-like activation mechanism. If true, then we would predict that complexes with lectin and serine protease activities would be found in the complement system of echinoderms. Currently, only C3-like and Bf-like genes have been found. It is possible that the early echinoderms had a lectin pathway then lost it, but this is unlikely. The ancient lectin pathway need not contain proteins as complex as the MBP/MASP complex. Those may have appeared later, before the urochordates and vertebrates split. In the coming years, if no putative lectin activator is found, then that could be considered evidence against this model. If that happened, then any model that does not require lectins as the initial activator will be supported.
While much of this model is speculative, it is based on the comparison of the functions and genetic sequences of complement proteins in different organisms. There are still many gaps to be filled in, but the foundation of the evolutionary pathway is clear. Much of the complement system was formed through a series of gene duplication events, co-opting the existing genes to add additional layers to this proteolytic cascade. This is not an uncommon idea in evolutionary biology, and there is much evidence to suggest that many serine protease cascades were generated through similar means (Krem and Di Cera 2002). While one could demand more evidence to support this story, it’s a much better story than what we could have come up with five years ago, which was much better than ten years ago, and so on. There is a trend of increasing information and knowledge that makes the story more precise, with fewer holes. To seek an answer to a particular gap in our knowledge is what science is all about. To doubt that the gap can be filled is unjustified. To claim that the existence of a gap is evidence that it cannot be filled is absurd.
The mammalian immune system is indeed a complex system. It may very well be irreducibly complex, as Behe suggests. However, complexity by itself does not eliminate evolution. By now it should be clear to the reader that in each system discussed in Chapter 6 of Darwin’s Black Box, there are models for their origins and ample evidence to support those models. He fails to acknowledge the strength of evidence supporting evolutionary models, or dismisses them without justification. Behe makes critical errors that distort his analysis. Let’s review the systems described by Behe and their supposed "evolvability":
Clonal Selection – Behe lists three components as part of the clonal selection system’s "core": the membrane-bound immunoglobulin, the signal transduction machinery (messenger), and the secreted Ig. According to him, none of these components can serve any function without the others. Behe is incorrect. The components of the signal transduction pathway exist in animals, such as insects, that do not possess the clonal selection system. Numerous innate receptors exist that induce a signal, but do not switch from a membrane-bound form to a secreted form upon activation. There are receptors that are expressed in membrane-bound form and secreted form, but are not induced to do so. Lastly, there exists a receptor with extreme homology to immunoglobulins and uses very similar signaling proteins, but does not switch forms. We can observe in nature functional systems that lack one or more of Behe’s "core" components. Whether or not those components are essential for function in the clonal selection system has nothing to do with how that system arose.
V(D)J Recombination – The "core" of the V(D)J recombination system consists of the antigen receptor genes, the recombination machinery, and the recombination signal sequences (RSSs). While a non-rearranging antigen receptor with clear homology to the immunoglobulins has yet to be discovered, there exist many innate receptors that can protect the host from invasion. When Behe states that "A primitive system with only one or a few antibody molecules would be …. not sufficient to make a difference", he is dead wrong. Insects, echinoderms, tunicates, lancelets, and agnathans have all been able to survive without the benefit of an adaptive immune system, relying on an array of innate receptors specific to common bacterial surface constituents. As for the recombination machinery, and the RSSs, Behe is correct to state that one is useless without the other. However, the proteins with homology to the RAGs, and sequences quite similar to the RSSs can be observed operating in bacterial and viral transposons. The current model that the adaptive immune system originated when a transposon inserted itself into an innate antigen receptor is alive and well. Since the publication of Darwin’s Black Box in 1996, this model has all but been confirmed by the discovery of transposase activity in the RAG proteins.
The complement system – Behe’s largest error in chapter 6 is choosing the complement system as an example of how evolution cannot produce irreducible complexity. He not only neglects to define which components form the "core", but lists nearly every protein involved in the pathway, suggesting that all are required for minimal function. He acknowledges only one of several functions of the complement system. He even implies that the system is dependent on antibodies. The discovery of subsets or precursors of the complement system in invertebrates lacking antibodies dispels that notion. The fact that these invertebrates’ complement systems do not have the one function he defined for it shows how faulty his IC logic is. However, whether or not the system is irreducibly complex is irrelevant. The evidence for its evolution is undeniable. There is a trend of increasing numbers of components of the complement system going up the tree of life. Arthropods have none, Echinoderms have a pair, tunicates have more, and so forth. At what point along this evolutionary route does Behe think that the barrier to evolution lays?
The errors made by Behe in Chapter 6 are not mere oversights. He has taken faulty assumptions, and made very strong conclusions based on them. Throughout the chapter Behe insinuates that no research is being conducted on the evolution of the immune system, and that no progress has been made. However, he is sorely mistaken. The literature cited in this review represents only a tiny subset of the total number of articles on the subject, and the total amount of research being conducted. Why then, does he state, "We can look high or we can look low, in books or in journals, but the result is the same. The scientific literature has no answers to the question of the origin of the immune system"? Although this statement is totally ludicrous, it would be unfair to suggest that Behe has intentionally misled readers into thinking that the scientific community is baffled by the immune system’s complexity. It would be fairer to say that he is merely ignorant of the abundance of research and articles that exist on the subject, or that he does not understand them. Behe is not an immunologist, and in the modern era of biology, it is nearly impossible to be an expert in all fields. Furthermore, Darwin’s Black Box is not a peer-reviewed article, but a book meant for the lay reader. Should he be expected to know every detail in the literature on immune system evolution? Absolutely. If Behe wants the reader to believe that he is an authority on the subject, then yes, he needs to know what every immunologist knows about the field. Is that unfair? Perhaps the inaccuracy is due to the difficulties in conveying a complex subject like the immune system to the lay reader. However, no immunologist would say that Behe’s analysis is accurate. The errors he makes display a lack of familiarity with the field. He should not have written a chapter on immune system evolution, nor made such strong conclusions. Either that or he should have sent the manuscript to an immunologist before publishing.
Since we cannot look at the genes of our ancestors, we must turn to our relatives. Armed with an understanding of how evolution works, we can probe through the genetic sequences of different organisms, and draw conclusions based on what is similar, and what is different. Similarities suggest homology, and by generating phylogenies of genes and families of genes from a large grouping of organisms, we can track when and how quickly genes diverged. We can piece together what steps occurred in the evolution of a system. We can use this information to generate models from which to make hypotheses, then test them in the lab. Based on those results, we can eliminate inaccurate models and fine-tune surviving ones. Ultimately, we’ll inch closer and closer to a more complete understanding of how the system works, and how it arose. For example, the model proposed for the origin of the complement system is based solely upon comparisons of different organisms. In time, more information on the origins of the other two systems will be revealed through comparative immunology, such as whether the immunoglobulin evolved from the T cell receptor or vice versa. It is startling that Behe would dismiss the use of comparative immunology, stating that it "does not address in molecular detail the question of how immune systems originated." Perhaps he should purchase a subscription to the journal Developmental and Comparative Immunology?
Behe is a biochemist, and prefers functional analysis over the comparison of gene sequences. The approach he proposes to analyze the "evolvability" of systems is a genetic one: Remove a component and see what happens. He feels that this method, through which irreducible complexity is identified, can reveal potential difficulties in the evolution of a system that scientists have previously overlooked. In doing so, irreducible complexity can potentially challenge the scientific community to think of new solutions in molecular evolution. But is his method useful? Does it actually reveal potential roadblocks? In each of the three systems described by Behe in chapter 6, I have neglected to discuss whether the systems are IC. Why? Because it doesn’t affect how the system could have evolved. Whether or not a particular component is required for function in the current system does not necessarily mean it was required in an ancestral system. C4 and C2 are required for complement activation in mammals, yet invertebrates do not possess them. The ability to switch from a membrane-bound form to a secreted form is essential for immunoglobulins, but not T cell receptors (TCR). As has been previously mentioned, Behe’s argument is essentially an argument from ignorance. He seems to require detailed knowledge of a system’s origin before he will consider the possibility that it evolved through "Darwinian" evolution. If the absence of C4 and C2 had never been discovered in invertebrates, would we conclude that the complement system could not have evolved? If TCRs were not present in jawed vertebrates, would we conclude that an antigen-receptor that was only membrane-bound could not serve a useful function? Behe’s method takes an "unevolvable until proven evolved" approach, which will invariably lead to false positives. Interestingly, the method used to disprove his inferences, comparative immunology, is dismissed by Behe. Perhaps the reason he dismisses it so quickly is because it can only serve to disprove him? The approach he proposes may be useful in determining the function of a system, and how each component contributes to it, but it is clearly useless in regards to origins.
The issue of peer-review, or lack thereof, is one that often arises in discussions of the ID movement. In fact, no critique of an ID article would be complete without mentioning it. No research article on ID has ever been published in a reputable peer-reviewed journal. ID advocates might suggest that the scientific community is biased against them, and rejects their articles for political reasons. Whatever the reason, ID advocates have instead opted for publishing popular books, which do not require the approval of the scientific community. Should we take these books seriously, since IDists would claim that they are being actively prevented from publishing in peer-reviewed journals? Individuals familiar with biology can judge the work for themselves, but lay readers cannot tell when they are being told good science, and when they are being misled. They cannot judge the accuracy of the science for themselves, at least not without great effort. Ultimately, there is a degree of trust that must be earned between author and reader. Trust that the information conveyed is accurate. Before a scientific journal will accept an article for publication, they send copies of the article to two or three scientists who are considered to be experts in the particular field encompassed by the article. They review the article with a fine-toothed comb, and point out any weaknesses and errors the authors may have made, and suggest experiments to strengthen it. The editor of the journal will decide whether or not to consider the article, and tell the author what improvements need to be made before they will accept it. Changes can be as subtle as changing a verb from "indicates" to "suggests". Only after the improvements have been made will the article be published. This system of peer-review ensures the high quality of research presented by articles in the journal, and prevents gross errors from being published. Darwin’s Black Box
does not present any original research, but is a review of the existing literature with a focus on origins. However, even though the research and knowledge presented has been published by others in peer-reviewed journals, the accuracy of Behe’s analysis is so poor that it clearly would not have passed peer-review had it been submitted to a journal. That is why peer-review is so important to scientific progress, and why the issue is brought up time after time by ID critics. Peer-reviewed journals stake their credibility on the articles they publish. They have earned the respect and trust of their readership.Skepticism and motivation
Oftentimes in the evolution/creation debate, the question of motivation arises. Proponents of evolution will state that the ID advocates are religiously motivated, and the ID advocates will in turn state that the evolution proponents are materialists, or dogmatic Darwinists. This form of ad hominem argues against the motivations of the individual, and distracts from the heart of their arguments. This is generally considered a fallacy, and should be avoided. However, much of the evidence of evolution is subject to interpretation, and often requires the experience of an expert to understand. If the lay person is unable to interpret the data for themselves, they must rely on the opinion of the expert. In that case, it may be important to understand the motivations and biases of those who claim to be experts on the subject. In chapter 6, Behe writes as though he considers himself an authority on immunology. However, many of his statements display a lack of understanding of some fundamental concepts of immunology. For example, he states that a non-rearranging antigen receptor would be useless to the organism. As has been previously mentioned, this is in stark contrast to what is observed. Behe also questions the validity of certain molecular mechanisms for mediating evolutionary change, such as gene duplication. He has demanded in each chapter detailed molecular explanations for how these systems could have originated. Usually, when an individual shows extreme skepticism for one concept, it is because they are already convinced by an alternative concept. In Behe’s case, that would be Intelligent Design. However, his degree of skepticism toward "Darwinian" evolution is not equally directed towards his own thesis. He has offered no detailed mechanisms for how these systems could have been designed, how they were implemented, and how often they occurred. In the 6 years following Darwin’s Black Box’s publication, he has made no attempt to answer any of these questions, nor conducted any research to further ID as a scientific concept. On the other hand, much progress has been made towards increasing our understanding of how these systems arose through evolution. In fact, about two-thirds of the articles cited here were published after 1996. So why does Behe continue to espouse ID? Why does he display such critical skepticism for evolution, when ID has nothing to offer? Why does he make such strong conclusions based on erroneous assumptions and an argument from ignorance? It is here that the issue of motivation arises, not as a justification for dismissing his arguments, but as an explanation for why he would continue to deny the abundance of evidence that supports an origin of these systems through "Darwinian" evolution.
If Dr. Behe reads this article, it would probably be unwise for him to respond personally, as it would serve him no useful purpose and just expose him to even more criticism. However, there may be others who agree with his arguments and would like to respond. To those persons I would ask that their response focus on the arguments specifically relating to immunology, such as those presented in the body of the article, and not the more generic arguments contained within the introduction and conclusion. Arguing from incredulity should be avoided, especially in the absence of a competing hypothesis. If the responder feels that a competing hypothesis exists, please present it in as much detail as possible, or refer to an article or book that contains the hypothesis. I would also request that any attempt to respond to this article address the following concerns:
The definition of irreducible complexity is too vague to be useful. Terms like "system", "function", "parts", "well-matched", and "interacting" need to be clearly defined.
The T cell receptor is a rearranging antigen receptor that lacks the ability to switch forms. Could the immunoglobulins have evolved from T cell receptors?
Innate, non-rearranging receptors are beneficial to the host. Behe is wrong to assert that "a primitive system with one or a few antibody molecules would be…not sufficient to make a difference).
The similarities between the V(D)J recombination machinery and transposases are not trivial. What explanation does Behe have for why the RAG proteins are able to catalyze transposition reactions?
Behe did not list the essential parts for the complement system. Additionally, he does not state what the function of the complement system is. What are the essential parts? What is the function?
How does the discovery of complement proteins in invertebrates affect Behe’s IC inference? Are antibodies required for complement activation? If not, then are the components specific to the classical pathway essential, as is suggested in chapter 6?
Behe often demands "quantitative calculations" for evolutionary models. What quantitative calculations can be offered in favor of ID?
What positive predictions can be made based on irreducible complexity?
What research has been conducted on ID since Darwin’s Black Box’s publication? What research has even been proposed?
Michael Behe presents a very bleak picture of how evolution could have produced the systems described in chapter 6 of Darwin’s Black Box. He makes several erroneous assumptions in his analysis, some of which demonstrate a lack of understanding of immunology. He only briefly mentions the current models, then dismisses them without justification. He demands detailed explanations, but does not offer any detailed alternatives in return. He scrutinizes evolutionary theory, then bases his IC to ID inference on negative evidence and arguments from ignorance. He makes no predictions, no hypotheses, and proposes no research. Darwin’s Black Box, and the concept of irreducible complexity is essentially just a giant argument from ignorance and a scientific dead end. Behe needs to be reminded of the old adage: "Absence of evidence is not evidence of absence".
Thanks to Andrea Bottaro, Nick Matzke, and the TD crew
Agrawal, A., Q. M. Eastman, et al. (1998). "Transposition mediated by RAG1 and RAG2 and its implications for the evolution of the immune system." Nature (London) 394(6695): 744-751.
Al-Sharif, W. Z., J. O. Sunyer, et al. (1998). "Sea Urchin Coelomocytes Specifically Express a Homologue of the Complement Component C3." J Immunol 160(6): 2983-2997.
Armstrong, P. B., M. T. Armstrong, et al. (1993). "Involvement of alpha-macroglobulin and C-reactive protein in a complement-like hemolytic system in the arthropod, Limulus polyphemus." Molecular Immunology 30(10): 929-934.
Armstrong, P. B. and J. P. Quigley (1999). "alpha2-Macroglobulin: An evolutionarily conserved arm of the innate immune system." Developmental & Comparative Immunology 23(4-5): 375-390.
Bartl, S., D. Baltimore, et al. (1994). "Molecular Evolution of the Vertebrate Immune System." PNAS 91(23): 10769-10770.
Behe, M. J. (1996). Darwin's black box : the biochemical challenge to evolution. New York, Free Press.
Clow, L. A., P. S. Gross, et al. (2000). "Expression of SpC3, the sea urchin complement component, in response to lipopolysaccharide." Immunogenetics 51(12): 1021-1033.
Daffre, S. and I. Faye (1997). "Lipopolysaccharide interaction with hemolin, an insect member of the Ig-superfamily." FEBS Letters 408(2): 127-130.
dos Remedios, N. J., P. A. Ramsland, et al. (1999). "Identification of a homologue of CD59 in a cyclostome: Implications for the evolutionary development of the complement system." Developmental and Comparative Immunology 23(1): 1-14.
Dreyfus, D. H. (1992). "Evidence suggesting an evolutionary relationship between transposable elements and immune system recombination sequences." Molecular Immunology 29(6): 807-810.
Endo, Y., M. Takahashi, et al. (1998). "Two lineages of MASP/C1r/C1s family in vertebrate." Molecular Immunology 35(6-7): 349.
Fenton, M. J. and D. T. Golenbock (1998). "LPS-binding proteins and receptors." Journal of Leukocyte Biology 64(1): 25-32.
Gokudan, S., T. Muta, et al. (1999). "Horseshoe crab acetyl group-recognizing lectins involved in innate immunity are structurally related to fibrinogen." PNAS 96(18): 10086-10091.
Hall, R. M., C. M. Collis, et al. (1999). Mobile gene cassettes and integrons in evolution.
Hiom, K., M. Melek, et al. (1998). "DNA transposition by the RAG1 and RAG2 proteins: A possible source of oncogenic translocations." Cell 94(4): 463-470.
Ishikawa, G., K. Azumi, et al. (2000). "Involvement of tyrosine kinase and phosphatidylinositol 3-kinase in phagocytosis by ascidian hemocytes." Comparative Biochemistry and Physiology Part A Molecular & Integrative Physiology 125A(3): 351-357.
Janeway, C. (2001). Immunobiology 5 : the immune system in health and disease. New York, Garland Pub.
Ji, X., K. Azumi, et al. (1998). "Opsonic complement C3 in the solitary ascidian, Halocynthia roretzi." Molecular Immunology 35(6-7): 363.
Ji, X., K. Azumi, et al. (1997). "Ancient origin of the complement lectin pathway revealed by molecular cloning of mannan binding protein-associated serine protease from a urochordate, the Japanese ascidian, Halocynthia roretzi." PNAS 94(12): 6340-6345.
Kairies, N., H.-G. Beisel, et al. (2001). "The 2.0-A crystal structure of tachylectin 5A provides evidence for the common origin of the innate immunity and the blood coagulation systems." PNAS 98(24): 13519-13524.
Kennedy, A. K., A. Guhathakurta, et al. (1998). "Tn10 transposition via a DNA hairpin intermediate." Cell 95(1): 125-134.
Krem, M. M. and E. Di Cera (2002). "Evolution of enzyme cascades from embryonic development to blood coagulation." Trends in Biochemical Sciences 27(2): 67-74.
Levashina, E. A., L. F. Moita, et al. (2000). "Conserved role of a complement-like protein in phagocytosis revealed by dsRNA knockout in cultured cells of the mosquito, Anopheles gambiae." Cell 104(5): 709-718.
Lewis, S. M. (1999). Evolution of immunoglobulin and T-cell receptor gene assembly.
Lewis, S. M. and G. E. Wu (1997). "The origins of V(D)J recombination." Cell 88(2): 159-162.
Lindstrom-Dinnetz, I., S.-C. Sun, et al. (1995). "Structure and expression of Hemolin, an insect member of the immunoglobulin gene superfamily." European Journal of Biochemistry 230(3): 920-925.
Matsushita, M., Y. Endo, et al. (2000). "Ficolins activate the lectin complement pathway." Immunopharmacology 49(1-2): 79.
Matsushita, M., Y. Endo, et al. (1998). "Complement-related serine proteases in tunicates and vertebrates." Current Opinion in Immunology 10(1): 29-35.
Matsushita, M. and T. Fujita (2001). "Ficolins and the lectin complement pathway." Immunological Reviews 180: 78-85.
Medzhitov, R. and C. A. Janeway (1997). "Innate immunity: The virtues of a nonclonal system of recognition." Cell 91(3): 295-298.
Melchior, R., J. P. Quigley, et al. (1995). "Alpha-2-Macroglobulin-mediated Clearance of Proteases from the Plasma of the American Horseshoe Crab, Limulus polyphemus." Journal of Biological Chemistry 270(22): 13496-13502.
Muta, T. and S. Iwanaga (1996). "The role of hemolymph coagulation in innate immunity." Current Opinion in Immunology 8(1): 41-47.
Nair, S. V., S. Pearce, et al. (2000). "A collectin-like protein from tunicates." Comparative Biochemistry and Physiology B 125B(2): 279-289.
Newton, R. A., D. A. Raftos, et al. (1994). "Chemotactic responses of hagfish (Vertebrata, Agnatha) leucocytes." Developmental and Comparative Immunology 18(4): 295-303.
Nonaka, M. (2001). "Evolution of the complement system." Current Opinion in Immunology 13(1): 69-73.
Nonaka, M., K. Azumi, et al. (1999). "Opsonic Complement Component C3 in the Solitary Ascidian, Halocynthia roretzi." J Immunol 162(1): 387-391.
Nonaka, M., T. Fujii, et al. (1984). "Purification of a lamprey [Lampetra japonica] complement protein homologous to complement C3 of the mammalian complement system." Journal of Immunology 133(6): 3242-3249.
Nonaka, M. and S. Miyazawa (2002). "Evolution of the initiating enzymes of the complement system." Genome Biol 3(1): REVIEWS1001.
Nonaka, M. and S. L. Smith (2000). "Complement system of bony and cartilaginous fish." Fish & Shellfish Immunology 10(3): 215-228.
Nonaka, M. and M. Takahashi (1992). "Complete complementary DNA sequence of the third component of complement of lamprey: Implications for the evolution of thioester containing proteins." Journal of Immunology 148(10): 3290-3295.
Nonaka, M., M. Takahashi, et al. (1994). "Molecular cloning of a lamprey homologue of the mammalian MHC class III gene, complement factor B." Journal of Immunology 152(5): 2263-2269.
Oettinger, M. A., D. G. Schatz, et al. (1990). "RAG-1 and RAG-2, adjacent genes that synergistically activate V(D)J recombination." Science (Washington D C) 248(4962): 1517-1523.
Roth, D. B., J. P. Menetski, et al. (1992). "Vdj Recombination Broken Dna Molecules with Covalently Sealed Hairpin Coding Ends in Scid Mouse Thymocytes." Cell 70(6): 983-991.
Sahu, A. and J. D. Lambris (2001). "Structure and biology of complement protein C3, a connecting link between innate and acquired immunity." Immunological Reviews 180: 35-48.
Sakano, H., K. Huppi, et al. (1979). "Sequences at the somatic recombination sites of immunoglobulin light-chain genes." Nature 280(5720): 288-94.
Schatz, D. G. (1999). "Transposition mediated by RAG1 and RAG2 and the evolution of the adaptive immune system." Immunologic Research 19(2-3): 169-182.
Shintani, S., J. Terzic, et al. (2000). "Do lampreys have lymphocytes? The SPI evidence." Developmental & Comparative Immunology 24(Supplement 1): S104.
Smith, L. C., K. Azumi, et al. (1999). "Complement systems in invertebrates. The ancient alternative and lectin pathways." Immunopharmacology 42(1-3): 107-120.
Smith, L. C., L. A. Clow, et al. (2001). "The ancestral complement system in sea urchins." Immunol Rev 180: 16-34.
Smith, L. C., C.-S. Shih, et al. (1998). "Coelomocytes Express SpBf, a Homologue of Factor B, the Second Component in the Sea Urchin Complement System." J Immunol 161(12): 6784-6793.
Smith, S. L. (1998). "Shark complement: An assessment." Immunological Reviews 166(0): 67-78.
Sottrup-Jensen, L., T. M. Stepanik, et al. (1985). "Common evolutionary origin of .alpha.2-macroglobulin and complement components C3 and C4." Proceedings of the National Academy of Sciences of the United States of America 82(1): 9-13.
Spanopoulou, E., F. Zaitseva, et al. (1996). "The homeodomain region of Rag-1 reveals the parallel mechanisms of bacterial and V(D)J recombination." Cell 87(2): 263-276.
Su, X.-D., L. N. Gastinel, et al. (1998). "Crystal structure of hemolin: A horseshore shape with implications for homophilic adhesion." Science (Washington D C) 281(5379): 991-995.
Sun, S. C., I. Lindstrom, et al. (1990). "Hemolin: An insect-immune protein belonging to the immunoglobulin superfamily." Science (Washington D C) 250(4988): 1729-1732.
Suzuki, M. M., N. Satoh, et al. (2002). "C6-like and C3-like molecules from the cephalochordate, amphioxus, suggest a cytolytic complement system in invertebrates." Journal of Molecular Evolution 54(5): 671-679.
Swarnakar, S., R. Asokan, et al. (2000). "Binding of alpha2-macroglobulin and limulin: Regulation of the plasma haemolytic system of the American horseshoe crab, Limulus." Biochemical Journal 347(3): 679-685.
Takahashi, H., G. Ishikawa, et al. (1997). "Cloning and tyrosine phosphorylation of a novel invertebrate immunocyte protein containing immunoreceptor tyrosine-based activation motifs." Journal of Biological Chemistry 272(51): 32006-32010.
Thompson, C. B. (1995). "New insights into V(D)J recombination and its role in the evolution of the immune system." Immunity 3(5): 531-9.
Thornhill, R. H. and D. W. Ussery (2000). "A classification of possible routes of Darwinian evolution." Journal of Theoretical Biology. 203(2): 111-116.
Van Gent, D. C., K. Mizuuchi, et al. (1996). "Similarities between initiation of V(D)J recombination and retroviral integration." Science (Washington D C) 271(5255): 1592-1594.
Xu, Y., S. V. L. Narayana, et al. (2001). "Structural biology of the alternative pathway convertase." Immunological Reviews 180: 123-135.
i One paper, "A classification of possible routes of Darwinian evolution", by Thornhill and Ussery, defines Darwinian evolution as, "Descent of organisms in which the following criteria are met: (i) intergenerational differences are very much smaller than inter-specific ones; (ii) no intervention by conscious agent(s) occurs; (iii) the frequency of mutations or other heritable modifications is unrelated to functional utility; and (iv) selection is the sole means by which heritable modifications are accumulated to form functional structures" (Thornhill and Ussery 2000)
ii The Thornhill and Ussery paper does include a discussion of irreducible complexity. However, because the term is not mentioned in the title or abstract, it fails to appear on a PubMed search.
iii According to citation index at http://webofscience.com/
iv The PubMed citation index features a tool which lists articles closely related to a selected article. Web readers can click here to see the list, or see http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?db=PubMed for details.
v For additional information on arthropods, see: http://tolweb.org/tree?group=Arthropoda&contgroup=Animals
vi For additional information on echinoderms, see: http://tolweb.org/tree?group=Echinodermata&contgroup=Animals
vii For additional information on urochordates, see: http://www.personal.psu.edu/users/m/x/mxg276/bio110h.html
viii For additional information on cephalochordates, see: http://www.personal.psu.edu/users/m/x/mxg276/bio110h.html
ix For additional information on cyclostomes, see: http://tolweb.org/tree?group=Hyperoartia&contgroup=Vertebrata
x For additional information on cartilaginous fish, see: http://www.ucmp.berkeley.edu/vertebrates/basalfish/chondrintro.html